- The Biomedical Data guide is designed to connect you to Biomedical Research resources available via UM Libraries. This guide provides information about statistical tests, finding data sources, data visualization resources, file naming and structure, and electronic laboratory notebooks. You can also access the learning resources offered in my workshops, covering topics in R, SPSS, Tableau, REDCap, GraphPad Prism, and Meta-Analysis.
- This guide is constantly evolving, and contact me at thilani.samarakoon@miami.edu if you find a tool or learning resource that could be included here.

Thilani Samarakoon
- Biomedical Data Librarian
- thilani.samarakoon@miami.edu
- (305) 243-9126

Cameron Riopelle
- Director of Research Data & Open Scholarship
- criopelle@miami.edu
- (305) 284-1524
Data visualization is the visual presentation of data and an essential step in data analysis.
As the image below illustrates, a good visualization should be the intersection of data, function, and design.
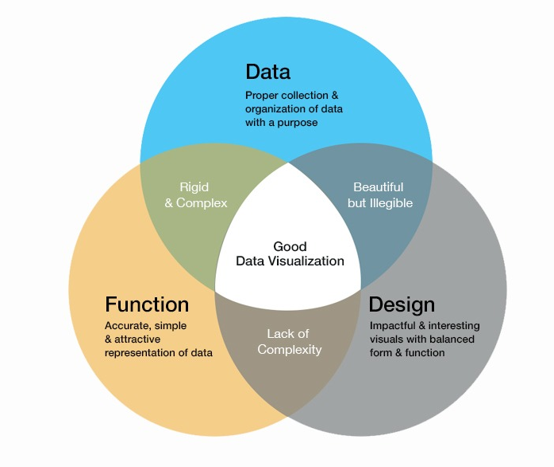
Visual elements like graphs, maps, tables, and plots help the audience identify trends, patterns, and distributions in the data. Data visualization is useful in converting complex data sets into meaningful information that the user quickly understands. Researchers in biomedical fields collect large amounts of data that are becoming increasingly complex.
- The Data Visualization Catalogue
- Better Data Visualizations: A Guide for Scholars, Researchers, and Wonks
- Designing Data Visualizations
- A Practitioner's Guide to Best Practices in Data Visualization
- Beginning Data Science in R: Data Analysis, Visualization, and Modelling for the Data Scientist
- Tasks, Techniques, and Tools for Genomic Data Visualization
- Know your audience
- Choose the right graph/table
How to choose the right graph for your data.
The data visualization catalogue.
- Declutter
Declutter-story telling with data.
Charts Do's and Don'ts.
- Focus attention
Practical Rules for Using Color in Charts
Choose Appropriate Visual Encodings
Tableau offers free one-year licenses to students. Click to request your license.
Tableau online resources are a great way to start learning the software.
R is a free software environment for creating visualizations. It requires some programming knowledge.
R offers additional packages to cater to any visualization task.
ggplot2: Creates custom plots. Refer to Cookbook for R to learn ggplot2.
GGally: Creates correlation matrix and survival plots.
gplots: Creates heat maps, Venn diagrams.
leaflet: Builds interactive maps.
Click to learn more about R packages.
Biomedical data visualization tools play a crucial role in transforming complex biological and medical datasets into visually interpretable formats.
Infographics create visual communications and provide an overview of the topic.
AHRQ Research Data Infographics
Picktochart
Canva
Cochrane Handbook for Systematic Reviews of Interventions, version 6.2
Meta-analyses are a subset of systematic reviews and play a fundamental role in evidence-based healthcare. It comes at the top of the evidence-based medicine pyramid.
- Combining studies in a meta-analysis can increase the overall statistical power to detect an effect.
- Help make a valid decision about intervention and identify the reasons for the variation.
- Settles controversies arising from apparently conflicting studies or evaluates new hypotheses.
- Search the published literature: Cochrane Library, Campbell Collaboration
- Registered as an ongoing review: Prospero, Cochrane
Eg: Study is restricted to a certain age group, symptoms experienced by the participants, location, written language
- Dichotomous variables (e.g., deaths, patients with at least one stroke)
- Count data (e.g., number of strokes)
- Continuous variables (e.g., blood pressure, pain score)
- Sample size
- Sensitivity, specificity
- Survival data
Statistical analysis is a two-step process.
- First stage: summary statistic is calculated for each study to describe the observed intervention effect.
- Second stage: A summary (combined) intervention effect estimate is calculated as a weighted average of the intervention effects estimated in the individual studies.
- Fixed effect model
- assumes all studies are estimating the same (common) treatment effect.
- Random effects model
- assumes observed estimates of treatment effect can vary across studies.
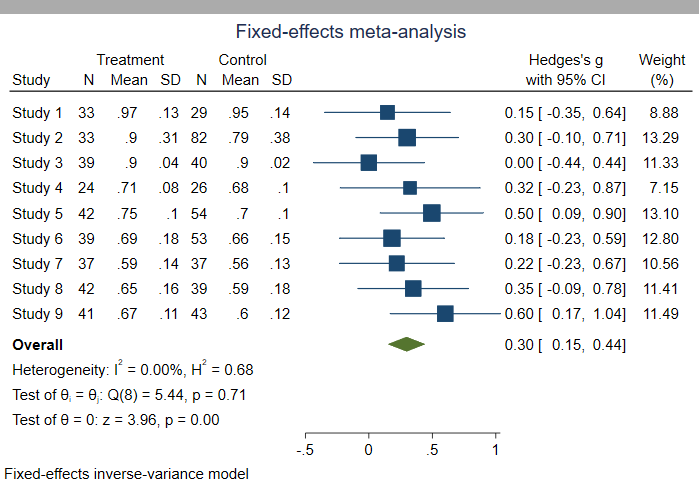
Source: Introduction to SAS. UCLA: Statistical Consulting Group.
- MEDLINE - Biomedical literature database
- CENTRAL - Cochrane Central Register of Controlled Trials
- CINAHL - Cumulative Index to Nursing and Allied Health Literature
- Embase - Biomedical and pharmacological (including MEDLINE titles)
- PsycInfo - Psychological, behavioral, social, and health sciences research
- ClinicalTrials.gov - database of clinical studies conducted around the world
- Web of Science, Scopus, Google Scholar
- Cochrane Interactive Learning: Calder medical library offers access to the online course.
- Revman training videos
- Cochrane Handbook for Systematic Reviews of Interventions: This is the official guide that describes in detail the process of preparing and maintaining Cochrane systematic reviews on the effects of healthcare interventions
- Systematic review resource guide at Calder medical library.
The site contains health-related data sets on various topics, including community health, medical devices, substance abuse, and hospitals. Data sets are organized by the topic. Some data sets are readily available, and in other cases, you will be directed to another site.
The site is home to more than 350,000 research studies conducted in more than 200 countries. The U.S. National Library of Medicine runs this resource.
The site offers a comprehensive guide on how to customize a search.
https://clinicaltrials.gov/ct2/help/how-find/index
NCHS is a part of the Centers for Disease Control & Prevention (CDC) and provides data and health statistics on crucial public health topics.
- Health, United States, is an annual report on trends in health statistics. It includes trend tables and figures on selected topics such as mortality, life expectancy, causes of death, and health care expenses, among other topics. https://www.cdc.gov/nchs/data/hus/hus18.pdf
- FastStats is a site within the NCHS that offers quick access to data, statistics, and reports on specific health topics. The topics are organized in alphabetic order and will direct you to more resources on the topic.
- The National Vital Statistics System provides access to most of the data on birth and deaths in the United States. There are links on the webpage that links to statistical reports, data sources, and data visualizations.
Cancer Query Systems - allows access to cancer data stored in an online database. The web-based interface is easy to use and retrieve statistics using a query-based system.
SEER*Stat - The data downloaded can be analyzed using the SEER* Prep and SEER*Stat statistical software.
Please refer to the webpage for installation instructions and tutorials.
NDA collects and stores data from and about human subjects across many research fields. It holds data collected by researchers funded by the National Institute of Alcoholism and Alcohol Abuse (NIAAA) and autism researchers. Also, it stores data from the Adolescent Brain and Cognitive Development Study, the Human Connectome projects, and the Osteoarthritis Initiative.
The website offers access to national, state, and regional data collected via the National Survey of Children’s Health (NSCH). The survey collects data on mental health, dental health, development, and many other child health indicators to improve child, youth, family, and community health and well-being. You can submit a data request to access the raw data or use the interactive query base system to search the data.
The survey collects data about the American public’s knowledge of, attitudes toward, and use of cancer- and health-related information. The data sets help researchers identify how adults use different communication channels to understand and obtain health-related information. HINTS data are free to download and analyze.
NTDBRDS is maintained by the American College of Surgeons and is the most extensive collection of U.S. trauma registry data. The researchers must submit an online request to obtain the data sets that are available at a cost.
HCUP's databases contain information on inpatient stays, emergency department visits, and ambulatory care.
Nationwide HCUP Databases
National Inpatient Sample (NIS)
- Kids’ Inpatient Database (KID) - KID Database Documentation
- Nationwide Ambulatory Surgery Sample (NASS) - NASS Database Documentation
- Nationwide Emergency Department Sample (NEDS) - NEDS Database Documentation
- Nationwide Readmissions Database (NRD) - NRD Database Documentation
State-specific HCUP Databases
- State Inpatient Databases (SID) - SID Database Documentation
- State Ambulatory Surgery and Services Databases (SASD) - SASD Database Documentation
- State Emergency Department Databases (SEDD) - SEDD Database Documentation
HCUPnet
HCUPnet is a part of the HCUP project and offers a free on-line query system to access data on inpatient stays (NIS, KID, SID, and NRD), emergency department visits (NEDS, SEDD, SID), and community (SID). You create an analysis set-up by clicking on “Create new analysis.” Once the analysis is completed, it can be customized using the “My analysis” menu.
The types of data/level of measurement mainly fall into two broad categories: continuous and categorical. By clicking on each test tab, you can learn more about the assumptions associated with the test and when to use it.
Parametric Tests
Continuous Data
- Example: Compare the means of two independent samples.
- Assumptions: Normal distribution (Shapiro-Wilk test), homogeneity of variances (Levene's test), independence of observations, no outliers.
- SPSS Demo
- Example: Compare the means of two dependent samples.
- Assumptions: Normal distribution (Shapiro-Wilk test), homogeneity of variances (Levene's test), independence of observations, no outliers
- Example: Compare independent sample means for more than 2 groups. Post Hoc - if equal of variances assumed select Tukey test. Otherwise, select Games-Howell.
- Assumptions: Normal distribution (Shapiro-Wilk test), homogeneity of variances (Levene's test) independence of observations, no outliers.
- Example: Compare means of 3 or more groups with the same participants.
- Assumptions: Normal distribution (Shapiro-Wilk test), homogeneity of variances (Levene's test) independence of observations, no outliers, sphericity (Mauchly’s test).
- Example: Examine the linear association and direction between two continuous variables. It takes a value between +1 to -1.
- Assumptions: Association between the two variables is linear, Normal distribution (Shapiro-Wilk test), no outliers.
- Example: Examine the linear association between a continuous outcome variable, and one or more independent variables.
- Assumptions: There should be a linear relationship between the 2 variables, no significant outliers, independence of observations, Normality.
Categorical Data
- Example: Examines the probability that an observation falls into one of two categories of a dichotomous dependent variable based on one or more independent variables that can be either continuous or categorical
- Assumptions: Dependent variable is dichotomous, independence of observations, linear relationship between any continuous independent variables and the logit transformation of the dependent variable.
Non-Parametric Tests
Continuous or ordinal Data
- Example: Compare the means of two independent samples when the normality assumption is violated. (alternative to an independent t-test)
- Assumptions: Independence of observations, distributions of the independent variable should have the same shape, independent variable consists of two categorical groups.
- Example: Compare the means of two dependent samples when the normality assumption is violated. (alternative to the dependent t-test)
- Assumptions: Independent variable consists of two categorical groups, dependent variable measured at the ordinal or continuous level, distributions of the independent variable should have the same shape.
- Example: Compare independent sample means for more than 2 groups when the normality assumption is violated. (alternate to ANOVA, an extension of Mann-Whitney U test)
- Assumptions: Independence of observations, independent variable consists of two or more categorical groups.
- Example: Compare means of 3 or more groups with the same participants when the assumptions of ANOVA with repeated measures are violated or the dependent variable measured is ordinal. (alternate to ANOVA with repeated measures)
- Assumptions: A group is measured 3 or more times, random sampling.
- Example: Examine the linear association and direction between continuous or ordinal variables when the assumptions have failed for a Pearson correlation. It takes a value between +1 to -1. (alternate to Pearson correlation)
- Assumptions: There is a monotonic relationship between the two variables.
- Example: Examine the association and direction between continuous or ordinal variables. Specially applied to small sample sizes and for variables with many ranks. It takes a value between +1 to -1. (alternate to Pearson correlation and Spearman correlation)
- Assumptions: There is a monotonic relationship between the two variables.
Categorical Data
- Example: Compare sample proportions to check if there is an association between the categories. A residual analysis will identify the specific cells that make the greatest contribution to a significant chi-square test.
- Assumptions: Independence of observations, expected cell count assumption (expected cell counts are all greater than 5)
- SPSS Demo
- Example: Compare sample proportions to check if there is an association between the categories. Used for small samples.
- Assumptions: Independence of observations, if the expected cell count is less than 5 for any cell, use Fisher’s exact test.
- SPSS Demo
- Example: Check the differences on a dichotomous dependent variable between two related groups.
- Assumptions: The dependent variables should be mutually exclusive.
Other commonly used tests:
- This method is used to study the reliability of the scale used in a survey (commonly used with Likert scale questions).
- The scale ranges from 0-1. If the alpha is 0.7 or above, the measure can be considered reliable. For example, a survey was designed with 10 questions to measure if the students are happy at school.
- The questions are on a Likert scale. The researcher can survey a sample and carry out the Cronbach analysis to estimate the reliability.
- This test is used to study the inter-rater reliability when two raters are evaluating a variable on a categorical scale.
- The scale ranges from 0-1. There is no clear cut-off for a good agreement. Normally 0.6-0.8 is considered good.
- There are a few assumptions to be met before conducting the test. The categories should be mutually exclusive, the raters are independent, and the same raters evaluate all the questions.
- This test is used to study the inter-rater reliability when two or more raters evaluate a variable on a categorical scale.
- If the raters are not fixed, meaning the same 2 raters are not evaluating the questions.
Resources:
- Statistical methods for the analysis of biomedical data, Woolson, Robert F.; Clarke, William R. (William Radue), New York: Wiley-Interscience, [2002]
- https://statistics.laerd.com/
- Parametric and Non-parametric tests for comparing two or more groups
- Choosing a statistical test
It is important that we understand the difference between repeatability, reproducibility, and replicability. Below is the terminology established by the Association for Computing Machinery.
Repeatability (Same team, same experimental setup)
- The measurement can be obtained with stated precision by the same team using the same measurement procedure, the same measuring system, under the same operating conditions, in the same location on multiple trials. For computational experiments, this means that a researcher can reliably repeat her own computation.
- The measurement can be obtained with stated precision by a different team using the same measurement procedure, the same measuring system, under the same operating conditions, in the same or a different location on multiple trials. For computational experiments, this means that an independent group can obtain the same result using the author's own artifacts.
- The measurement can be obtained with stated precision by a different team, a different measuring system, in a different location on multiple trials. For computational experiments, this means that an independent group can obtain the same result using artifacts which they develop completely independently.
Goodman et al. (2016) proposed methods reproducibility, results reproducibility, and Inferential reproducibility as broader categories for research reproducibility.
The NIHPG include the better performance of statistical analysis, transparency in reporting, data and material sharing, consideration of refutations, and consideration of establishing best practice
guidelines. These guidelines encourage authors to provide complete information about their studies for better reproducibility.
Citation of BioResources in journal Articles (CoBRA)
This provides a checklist for the citation of bioresources (biological samples, data, and databases) used in scientific journal articles.
Animal Research: Reporting of In Vivo Experiments (ARRIVE)
This is a checklist of recommendations to improve the reporting of research involving animals – maximizing the quality and reliability of published research, and enabling others to better scrutinize, evaluate and reproduce it.
The TOP Guidelines
Transparency and Opened Promotion (TOP) provides guidelines to enhance transparency in journal publications. The main categories include citation, data transparency, analytical methods (code) transparency, research materials transparency, preregistration of studies, preregistration of analysis plans, replication.
Equator network
Enhancing the QUAlity and Transparency Of health Research (EQUATOR) is a network is an international initiative that seeks to improve the reliability and value of published health research literature by promoting transparent and accurate reporting and wider use of robust reporting guidelines.
PRISMA
Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) is an evidence-based minimum set of items for reporting in systematic reviews and meta-analyses. PRISMA primarily focuses on the reporting of reviews evaluating the effects of interventions.
DRYAD: Best practices for creating reusable data
DRYAD is a nonprofit membership organization that is committed to making data available for research and educational reuse. These guidelines help make the datasets findable, accessible, interoperable, and reusable (FAIR).
Below are a few steps we can take to improve reproducibility in research.
- Provide appropriate citations for data and materials
- Post data and code in a trusted repository
- Design transparency
- Preregistration of studies
- Sharing data
- What does research reproducibility mean?
- Rethinking research reproducibility
- Reproducibility in Science: Improving the Standard for Basic and Preclinical Research
- Reproducibility vs. Replicability: A Brief History of a Confused Terminology
- Teaching Computational Reproducibility for Neuroimaging
- Six factors affecting reproducibility in life science research and how to handle them
- The Road to Reproducibility in Animal Research
- Reproducibility and replicability of systematic reviews
- Files should include only letters, numbers, and underscores/dashes.
- No special characters.
- No spaces; Use dashes, underscores, or camel case (like-this or likeThis)
- Use a YYYYMMDD format for dates.
- The file name should describe their content. Remember length can be a limitation here.
- Be consistent: Use the same naming schema for related files.
- Be unique with the names even if they are stored in different folders.
- Keep a record of the naming convention.
- Use ordinal numbers (v1,v2,v3) for major version changes and decimals for minor changes (v1.1, v2.6).
- Use leading zeros so that they sort in order (e.g., 001, 002, 003).
- Project or experiment name
- Lab name/location
- Name of the researcher
- Date of creation
- Content of the file
- Experimental condition
- Sample number
- Eg: 20201223_ProjectName_ExperimentName_Control_012
- Decide on the file structure before you start the project. Think about the project complexity, the number of collaborators, and the length of the project.
- Be consistent.
- Create a README file.
- Organize the files systematically within folders.
- Folder and file names should reflect the content to help researchers locate data faster.
- Keep your raw data and analyzed data separately.
- Project (project stages)
- Year
- Location (geographic location, project site)
- File Type
- Team members
- Experiments (instruments, experiment types)
- Biomedical Research Data Management at Harvard
- Online short-seminar: Organize your files
- File naming and organizing best practices
- Folder and file naming convention – 10 rules for best practice
- Data can be accessed and organized from anywhere, anytime.
- Easily searchable.
- Data can be shared with an individual or a group in multiple formats. This facilitates collaborations.
- Backup and copy data become more efficient.
- Secure: data cannot be misplaced or leave the premises.
- Data management requirements by funding agencies can be met with ELNs.
- PI can monitor data and documentation.
- Identify key features and requirements for your lab.
- Test out a few products before purchasing. Most of the vendor products offer free trial periods and training videos. This is an excellent way to explore the software and assess if it fits your laboratory requirements. Alternatively, you can talk to peers or other institutes using the software to get their feedback.
- Cost: There are Implementation fees, maintenance fees, and training fees associated with the purchase of the software.
- Security: Most of the ELNs provide compliance with FDA 21 CFR Part 11, which regulates electronic documentation and signatures. The ELNs allow the group to customize individual and group access permissions. Not all members of the group require the same access privileges.
- Meet the best practice of organizing, sharing, and archiving data
- Docollab: This is a collaborative workspace for projects, tasks, documents, spreadsheets, and diagrams. All the content is stored and transferred in encrypted form and Docollab's infrastructure undergoes continuous monitoring and auditing in compliance such as HIPPA Business Associate Amendment.
- LabArchives: This is a browser-based platform that is used to manage laboratory data and help to organize the workflow in a research laboratory. Its products include ELN for research, inventory, and scheduling.
- RSpace: This is an affordable and secure enterprise-grade Electronic Lab Notebook (ELN) that catalyzes research and allows organizations to enjoy the benefits, efficiencies, and long-term cost savings of centralized, paperless data storage, as well as lab management, IP protection, secure collaboration, sample tracking and integration with the institute’s existing archiving system.
- Cynote: This is a laboratory notebook using a version control system and independent date-time stamping (as notarization), in order to ensure record accountability, auditing, and conforming to US FDA 21 CFR 21's rule on electronic records.
- LabJ-ng: This is a laboratory notebook for organic chemists. It has a client-server architecture with a web browsers interface. Data is stored on MySQL database. Chemical reactions are drawn in the java applet window. See project website for example.
- MyLabBook: MyLabBook is a free and open-source electronic lab notebook (ELN) built on the Drupal v.8-9 CMS. The basic installation provides capabilities for collecting, displaying, and sorting data with a sample data set and can be modified for different experiments. It can easily accommodate hundreds of users with a role-based permission system.
- Electronic Lab Notebooks: What they are (and why you need one)
- Biomedical Research Data Management at Harvard
- Keeping a Laboratory Notebook
- NNLM guide on Electronic Laboratory Notebook
I conduct a workshop series in the Fall and Spring covering topics in R, SPSS, Tableau, REDCap, GraphPad Prism, and Meta-Analysis.
Multiple workshops are available in the Fall and Spring semesters.
Spring 2024
Register here
You can access the material covered in the workshops below. The tutorials provide step-by-step instructions to learn the software. After going through the content, please contact me at thilani.samarakoon@miami.edu for assistance if you have any further questions.
Training Resources:
- Introduction to Tableau (download link)
- Introduction to R (download link)
- Data visualization with R (download link)
- SPSS (download link)
- Meta-Analysis (download link)
- Introduction to REDCap (download link)
- Introduction to GraphPad Prism (download link)
Additional resources:
Data and Visualization Services Department offers additional workshops and training resources. Please check out the complete list here.

Thilani Samarakoon
- Biomedical Data Librarian
- thilani.samarakoon@miami.edu
- (305) 243-9126

Cameron Riopelle
- Director of Research Data & Open Scholarship
- criopelle@miami.edu
- (305) 284-1524